http://ex-ample.blogspot.fr/2012/05/list-of-software-molecular.html
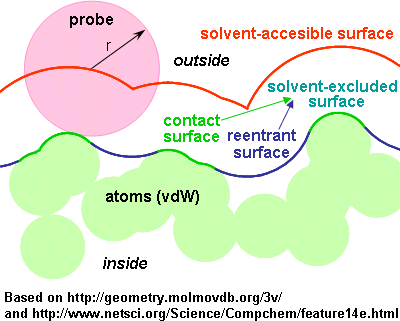
The solvent-accessible surface area (SASA) is the "virtual surface area" of a molecule that is "accessible" to a solvent.
SASA is typically calculated using the "rolling ball" algorithm [Shrake & Rupley,1973; Environment and exposure to solvent of protein atoms. Lysozyme and insulin. J Mol Biol 79(2):351-71. doi:10.1016/0022-2836(73)90011-9]. This algorithm uses a sphere (of solvent) of a particular radius to "probe" the surface of the molecule. A typical value is 1.2-1.4Å, which approximates the radius of a water molecule. Another factor that affects the results is the definition of the van der Waals radii of the atoms in the molecule under study. To generate the Van der Waals surface simply use zero (or 0.001) as the solvent radius.
The hydrogen atoms may be implicitly included in the atomic radii of the 'heavy' atoms. In addition, the number of points created on the van der Waals surface of each atom determines another aspect of discretization, where more points provide an increased level of detail.
[http://en.wikipedia.org/wiki/Accessible_surface_area]
The solvent-excluded surface (also known as the molecular surface or Connolly surface), is imagined as a cavity in bulk solvent (effectively the inverse of the solvent-accessible surface). It is also calculated in practice via a rolling-ball algorithm developed by Frederic Richards and independently implemented three-dimensionally by Michael Connolly in 1983 and Tim Richmond in 1984. Connolly spent several more years perfecting the method [ doi:10.1016/0263-7855(93)87010-3].
Try this stand-alone program: molekel, an open-source multi-platform molecular visualization program. http://molekel.cscs.ch/wiki/pmwiki.php, or Jmol...
The accessible surface is therefore sort of an expanded van der Waals surface.
It is larger and more external than the "molecular" surface.
How to obtain them in Jmol:
http://jmol.sourceforge.net/docs/surface/
Jmol, as Rasmol, uses a sphere with 1.2 Å radius for water; Chime uses 1.2 Å for dots and 1.4 Å for surfaces. Other programs/authors use 1.4 Å. The probe radius can be defined using the set radius command (Jmol).
-----------
example: Hemoblogin and H2O2: http://www.jbc.org/content/282/7/4894.full; Structural Basis of Peroxide-mediated Changes in Human Hemoglobin, A NOVEL OXIDATIVE PATHWAY; 2007; The Journal of Biological Chemistry, 282, 4894-4907.


No comments:
Post a Comment