Ref: http://www.caduser.com/reviews/reviews.asp?a_id=66
Healing the wounds of data conversion
From CAD User AEC Magazine Vol 13 No 03 - March 2000
Having, hopefully, encouraged much wider sharing of data and models throughout the CAD/CAM/CAE market, CAD User now sets out to draw your attention to common problems you will encounter when you do so.
There are two main suppliers of kernel software in the market, who provide the basic routines that are used by the overwhelming majority of application software packages. Spatial's ASIC is used to provide the guts for AutoCAD and one or two other packages, while Parasolid, developed by Unigraphics Solutions, is used by most mid-range CAD application developers, such Solid Edge, Solid Works, MicroStation Modeller, Top Solid, etc. Parasolid is also claimed to be the only kernel that is used in high-end 3D modelling systems. Parasolid has a greater share of 3D modelling software companies, due mainly to some advanced modelling capabilities that are particularly suited to solving the problems inherent in producing accurate and functional 3D models on screen.
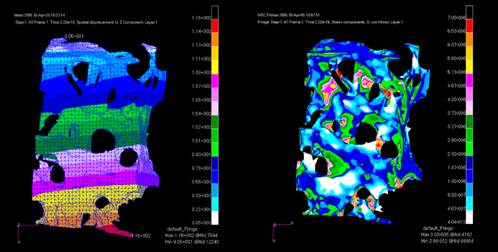
Design FEAts Besides enhancing visualisation, the ultimate aim in 3D modelling is to create digital representations of the putative finished product - and to use these to simulate the proposed function of the object (Working Model 4D) or to perform Finite Element Analysis, eliminating design flaws at the earliest possible stage. To do so, the digital geometry and topology of the object has to be correct - ie, boundaries must be viable, tangents must meet surfaces, surfaces must comply with recognised standards (STEP), and lines and thicknesses must be complete and accurate. Actually, accuracy is more important within finite element analysis than motion simulation - the latter is merely an extension of the cartoonist's craft, where a number of successive pages with slightly differing drawings are printed to the screen. Inconsistencies will occur when data is imported into one system from another, especially where that other system - possibly based on another kernel - is using different standards of accuracy,. Bad communications can also create anomalies in the data. Peter Kerwin, Parasolid's business development manager, calculates that up to 20 per cent of models imported into applications using its kernel software contain errors that have to be accommodated for before they can be used. Parasolid works to an accuracy of 10-8, in a world size of 103, giving an accuracy range of 1011, which is an order of magnitude above other kernel modellers (ACIS, for instance, can create models with a magnitude of 104, but has an accuracy of 10-6, resulting in an accuracy range of 1010). Unit sizes, are, of course, entirely arbitrary in each case, although Parasolid usually works to 1 metre and ACIS to 1mm. CAD developers using either of the kernels set their own units, of course, but I believe, like Parasolid, that 1 kilometer sized models (103 units) represent a reasonable maximum size of model for most purposes. Within the Parasolid kernel, there exists a series of functions that tolerate a certain amount of inaccuracy in imported data. This feature has been called Tolerant Modelling. It enables Parasolid to accept models with an accuracy of no greater than 10-4, closing gaps in boundaries, sorting out multiple intersections and eliminating spikes. More advanced eradication of anomalies, however, needs to be dealt with by a more powerful tool and Parasolid, therefore, has released a set of utilities, called PS/Bodyshop, that incorporates Tolerant Modelling capabilities, but also adds a number of other functions. PS/Bodyshop - the name is evocative of the makeover and titillation end of the motor Industry - kicks in from within Parasolid-based applications to manipulate data in a number of different ways. Closing adjacent faces and the edges on which they should lie is handled by Geometry Healing, the junctions between each being reconstructed. Tangential edges are healed using 3D Solver through Constraint Based Healing and Geometry Repair sorts out self-intersections and discontinuities in translated geometry. Short edges, sliver faces and duplicate geometry - all features that regularly crop up in imported data, especially older data where numerous additions and deletions have been made in drawings - can be removed and Trimming data, healed to create valid faces, can be successfully converted to solids. Should the user then wish to export the model outside Parasolid's world, then geometrical and topographical features that may prevent subsequent use are kindly removed. PS/Bodyshop is normally installed at the server end of the design group system and, as such, is generally transparent as a toolkit to the user. In operation, it highlights errors of concern, asking the user whether the data shown should be redefined or reconfigured.
PS/STEP, PS/IGES and PS/VDA-Fs Concurrent with its release of PS Bodyshop, Parasolid has brought out its own utilities that provide its users with access to industry standard representations of geometric and topographical data. IGES is an international neutral file format for CAD data. PS/IGES enables bi-directional translation of geometric data in IGES format in and out of other CAD systems. Parasolid has produced an API that enables PS/IGES to be incorporated within any Parasolid-based application. JAMA-IS, an IGES protocol developed by the Japanese motor industry, is supported by PS/IGES.
Translation classes Similarly, STEP is the ISO standard neutral file format for defining model surfaces and enabling the exchange of product model data. Using the AP203 protocol to describe the geometry and topology of CAD models, it has been so defined to allow the interchangeability of data between applications with different storage formats. PS/STEP translates data bi-directionally between Parasolid and STEP-AP203. It can also read STEP-AP203 files and create a model from them. Besides supporting STEP-AP203, PS/STEP also supports AP214 Class II, class III, class IV, class V and class VI entities. PS/VDA-FS translates files between the VDA Surface Data Format Interface format and Parasolid XT. VDA is a German standard for transferring surface data, developed by the German Automobile Manufacturers Association (VDA - Verband der Automobilindustrie). Parasolid API is used as a means of plugging in this translator toolkit as well. IGES is mainly a surface-based system, taking each space and creating a trimmed surface for it. The sheets are held in space relying on implied connectivity only - hence the need for them to be stitched together to create a solid model, using extensive human intervention. Both PS/IGES and PS/VDA-FS allow the user to stitch loose faces or sheets into solid models PS/IGES has numerous features that provide highly accurate translation and exchange of data. The transfer of data can be controlled by tunable parameters, whilst callback functions monitor conversion progress - allowing the conversion to be aborted at any stage. Reader modules check the syntax of IGES files, oversee conversion of IGES entities and monitor sewing of surfaces - and preserve entity, label, colour and layer attributes during translation. Root and subordinate nodes are identified and subsequently converted to Parasolid entities, which are then scaled to meters to conform to Parasolid scales of accuracy. STEP, on the other hand, stores the topology of the object, stating that edges may be co-incident. The data that creates the solid model is already there in B-Rep form (Boundary representation) and, knowing where the edges are, there is no need to stitch the surfaces together. PS/STEP incorporates Parasolid's body checking and tolerant modelling capabilities during translation.
Dual-kernel system A recent convert to Parasolid kernels is Visionary Design Systems, whose IronCAD, formally an ACIS-only system, has now been twinned with Parasolid to become the first dual-kernel system. IronCAD uses both kernels simultaneously, switching back and forth when needed. The principal benefit is obviously the ability to work on models developed under either kernel, even to the extent of combining data from either kernel into a single model. IronCAD enables users to switch from one kernel to the other when problems are encountered in one - say, complex bends - that can only be handled by the other. The nanosecond switch is usually invisible to the user. Parasolid is fundamentally involved in interoperability, a basic tenet of EDM. Although they supply a large number of application developers in the CAD/CAM/ CAE markets, each of these enhances and develops the kernel to suit its own particular needs and idiosyncrasies. This creates, along the way, the little sprites and elves that will cause mischief when users try to swap data with each other. As a principal supplier, therefore, they need to remain at the forefront of tool development, ensuring usability of data from whatever source, with or without their kernel. Parasolid was developed by Unigraphics Solutions and has become a line of business in its own right, developing, promoting and selling its kernel both to associated companies and their competitors. Unigraphics Solutions has Solid Edge and iMan under its wing as additional lines of business (not forgetting Unigraphics, the package, itself). Parasolid's user base in mid-range CAD and CAM, and in high-end CAD systems, includes a host of familiar names. CIMData says that 80 per cent of companies with PC CAM solutions are Parasolid licensees. How many people use Parasolid? According to CEO John Mazzola, the new millennium saw 440,000 active seats installed, heading rapidly towards the half million mark! Parasolid and ACIS each share about 40 to 45 per cent of the kernel solid modelling market, although Parasolid has more end user applications - Spatial's ACIS has a vast number of AutoCAD seats. Most of the ACIS seats are in the low-end 2D CAD market, whilst Parasolid has a higher share of the 3D market. Spatial is developing its interoperability and web-based capabilities, and has recently acquired InterData Access and Sven Technologies to assist them.
Standard arguments Until recently, one of the greatest curses of this industry was the total inability of suppliers to accept industry-wide standards. Everybody thought they had it right and, well, everybody else would eventually have to follow suit. It was also the only means of protecting investments in a very threatening environment. Now, with the ease of whirling data around the globe, comes the imperative for that data to be understood once received. A great deal of current emphasis in EDM/PDM software is placed on interoperability tools. However, core software is being developed by fewer outfits, as it requires vast amounts of resource to produce the highly advanced and specialist tools that are now demanded. Spatial and Parasolid provide the bones for many application software providers who subsequently tweak the core software to suit their own customers' needs. Underlying data structures and practices remain identical, though, boding well for future transparency of data. People will, in future, be able to work better and cheaper, without the need for time-consuming and costly manipulation of incoming data.
CU

 orbitals of unsubstituted cyclopentadienone. Using previously developed carbonylative coupling reactions, a series of tetraarylcyclopentadienones was synthesized, accessing a range of substituents not previously available. The UV-vis spectra of these molecules were compared to their calculated wave functions and predicted transitions. A quantitative structure-activity relationship was discovered that may greatly simplify prediction of band gaps for oligomers and polymers built from these tetraarylcyclopentadienones.
orbitals of unsubstituted cyclopentadienone. Using previously developed carbonylative coupling reactions, a series of tetraarylcyclopentadienones was synthesized, accessing a range of substituents not previously available. The UV-vis spectra of these molecules were compared to their calculated wave functions and predicted transitions. A quantitative structure-activity relationship was discovered that may greatly simplify prediction of band gaps for oligomers and polymers built from these tetraarylcyclopentadienones.