code: http://spidergl.org/js/spidergl.js
http://sourceforge.net/projects/spidergl/
GALLERY:
http://spidergl.org/code.php
1/ Web application, a shader authoring tool:
http://spidergl.org/meshade/index.htmlcode generation
(input just a mesh, file= .obj)
VERY GOOD.
Try it online or download the zipped archive (an HTTP server is needed):
2/ Visualization Methods for Molecular Studies on the Web Platform
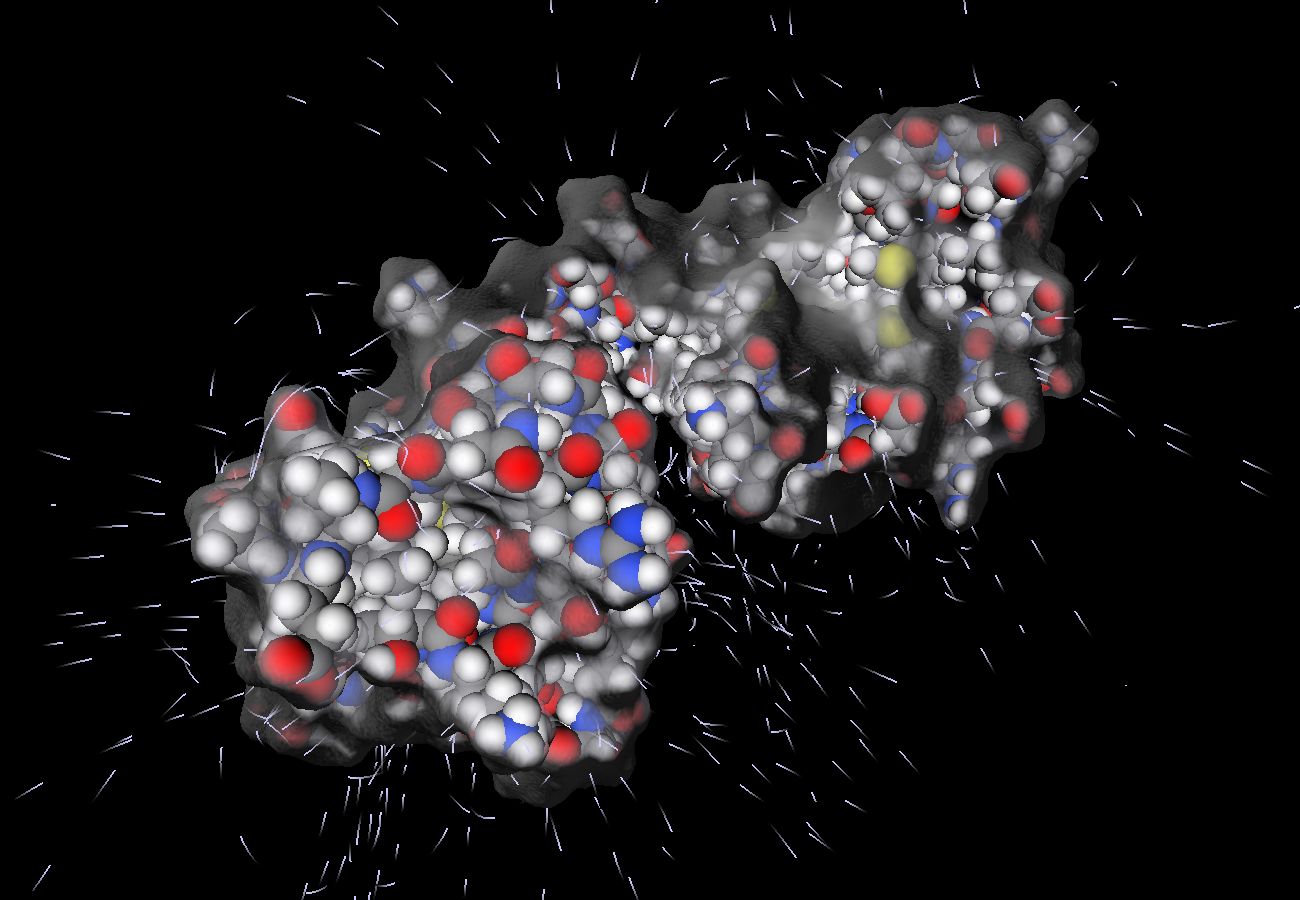
http://vcg.isti.cnr.it/Publications/2010/CADZS10/Standard representation of molecular surface properties using color ramps and field lines (leftmost), the same properties drawn using complex shading techniques (center) and the electrical interaction of two proteins (rightmost), rendered on a Web Page by using SpiderGL and WebGL.
Molecular visualization Off-Line and On-Line
The solution of the 3D structure of myoglobin in 1958 by Kendrew [Kendrew et al. 1958] marked the beginning of the new era of protein structural biology. Since then, a large number of protein structures have been solved and today the Protein Data Bank counts over 60.000 entries [Berman et al. 2003]. With the availability of all these data and the advance of computer graphics technologies, many research groups have developed tools for the manipulation and visualization of 3D structures such as VMD [Humphreyet al. 1996], SPDBViewer [Guex and Peitsch 1997], Chimera [Pettersen et al. 2004] and PyMOL [Delano 2002]. Beside working on the atomic structure, most programs can nowadays also calculate surface features such as electrostatic potential (using, for example, tools like APBS [Baker et al. 2001] or DelPhi [Rocchia et al. 2002]) and hydropathy [Kyte and Doolittle 1982].
In addition to the many standalone visualization tools, there are also web viewers especially designed for molecular structures, such as Jmol [jmo 2002] and MDL Chime, which represent a simple way
to visualize molecules directly on browser. MDL Chime, used by the Protein Explorer website was gradually phased out in favor of Jmol, which is nowadays the most used plugin for molecular visualization, used by websites such as Proteopedia and RCSB PDB Protein Data Bank.
Following the advance of techniques for the generation of CG movies, in the last few years many different groups focused on the creation of animated movies depicting biological molecules and
cellular processes. The movies range from the simple representations of the mechanical functioning of a single protein, to complex events involving many subjects. These works are important scientific efforts and add to their educational value the bonus of rising interest in the general public to approach biology. Some of these examples are collected on websites [McGill 2010; SCIVIS 2005].
3D Content on Web
The Virtual Markup Modeling Language (VRML) [Raggett 1994] (then replaced by X3D [Don Brutzmann 2007]) was proposed as a text based format for specifying 3D scenes in terms of geometry and material properties and for the definition of basic user interaction. The format itself was a standard, but the rendering in the web browser was relaying on specific plugins. The Java Applets are probably the most used method to add dynamic content, not necessarily 3D, in the web browsers. The
philosophy of Java applets is that the URL to the applet and its data are put in the HTML page and then executed by the Java Virtual Machine, a third part component. The implementation of JVM
on all the operating systems made Java applets ubiquitous and the introduction of binding to OpenGL such as JOGL [JOG ] added control on the 3D graphics hardware. A similar idea lies behind the
ActiveX [Microsoft Corporation 1996] technology, developed by Microsoft from 1996. Unlike Java Applets, ActiveX controls are not bytecode but dynamic linked Windows libraries which share
the same memory space as the calling process (i.e. the browser), and so much faster to execute. These technologies enable the incorporation of 3D graphics in a web page but they all do it by handling
a special element of the page itself with a third party component.
WebGL [Group 2009b] is an API specification produced by the Khronos group [Group 2009a] and, as the name suggests, defines the JavaScript analogous of the OpenGL API for C++. WebGL
closely matches OpenGLjES 2.0 and, extremely important, uses GLSL as the language for shader programs, which means that the shader core of existent applications can be reused for their
JavaScript/WebGL version. Since WebGL is a specification, it is up to the web browsers developer to implement it. At the time of this writing, WebGL is supported in the nightly build versions of the most used web browsers (Firefox, Chrome, Safari), and a number of JavaScript libraries are being developed to provide higher level functionalities to create 3D graphics applications. For example WebGLU [DeLillo 2009], which is the WebGL correspondent of GLU [OpenGL ARB ], provides wrappings for placing the camera in the scene or for creating simple geometric primitives, other libraries such as GLGE [Brunt 2010] or SceneJS [Kay 2009] uses WebGL for implementing a scene graph based rendering and animation engines.
Implementation spiderGL
Looking at a webpage with dynamic SpiderGL content, it is possible to see that all of the page logic is defined in the scripting part of the HEAD section, while on the BODY section there is just the page
structure and the interface elements that will be used for user interaction (like buttons, text areas and other controls). Among these elements, the most important is an html canvas object, that is the place where the WebGL layer does the on-screen rendering. ("< " are transformed : "[")
[canvas id="SGL_CANVAS" style="border: 1px solid gray" width="900" height="600"][/canvas]
This canvas is registered as the output area at the end of the scripting; a specific function connects the various events of the canvas to a script object.
v a r glMolViewer = new SpiderGLMolViewer ( ) ;
sglRegisterCanvas("SGL_CANVAS" , glMolViewer , 3 0 . 0 ) ;
The glMolViewer object is the main actor for the scene setup and rendering of our molecular visualization. The structure of this object employs the event handling subsystem provided by SpiderGL, which is inspired from the one used by the GLUT library [Kilgard ]. Each event coming from the canvas triggers a specific function with a given name and parameters; SpiderGL exploits the JavaScript language feature to give the possibility to dynamically add or remove listeners and redirect events. In this simple example, the only listener is the main object itself. (...)
This development process is straightforward for someone with an experience in graphical programming, while may prove to be difficult for users with a different background, like biology, physics or chemistry. This kind of setup is for sure more difficult to master with respect to setup of other existing platforms, like Jmol which, true to their nature, provide much simpler (but restrictive) access to their scene graph, with specific functions to import data and a series of predefined rendering modes. However, the gain in terms of flexibility and expressive power vastly compensate the initial steeper learning curve. Moreover, the learning of this technology is made easier by the possibility of initially use the higher level structures and functions implemented by SpiderGL to easily setup a basic visualization scheme and then start playing with lower level functions to obtain more complex effects. It is also important to note that most of the available JavaScript utility/UI libraries on the net may be used in conjunction with SpiderGL, adding more ready-made components to assemble a powerful, interactive, webpage.
3/ landscape refelction/refraction on a cup:
http://spidergl.org/example.php?id=12This example uses how to use cube maps and spherical harmonics to render objects with a natural light effect. GOOD.
4/techno
SpiderGL: A JavaScript 3D Graphics Library for Next-Generation WWW


No comments:
Post a Comment